Expanding the capabilities of targeted integration
Knock-in of a 50 kb construct into human iPSCs using TARGATT™ technology
Precise integration of large DNA payloads into mammalian genomes remains a fundamental challenge in cell line engineering, whether for basic research or therapeutic development. While CRISPR-based methods excel at knock-outs and point mutations, they struggle with insertions longer than a few kilobases due to the limitations of homology-directed repair and are known to generate off-target mutations¹. Lentiviral systems can deliver larger cargos but integrate randomly and often in multiple copies, while transposon-based methods similarly lack site specificity². These limitations have constrained applications requiring large, complex genome insertions, including screening libraries for variant assessment, multi-protein complex expression, stable packaging line generation for gene therapy, CAR-T cell engineering with logic gates, and humanized disease models³.
Accelerating complex genome engineering with rapid synthesis of ultra-long DNA constructs
Site-specific recombinases offer an attractive and proven alternative for large cargo integration. Applied StemCell’s TARGATT™ technology utilizes serine integrases and the mammalian H11 safe harbor locus to achieve efficient, site-specific integration and reliable gene expression⁴. The system has been previously validated for payloads up to 20 kb with integration efficiencies exceeding 40% in iPSCs without selection⁴, although the upper size limits for TARGATT™ cargo capacity have not been fully explored.
Here we demonstrate the successful integration of a 50 kb multi-protein construct into human iPSCs using TARGATT™ technology, enabled by Ansa Biotechnologies’ rapid enzymatic DNA synthesis platform. The construct, synthesized in less than one month — compared to typical two- to six-month timelines for constructs of this size — contains fluorescent reporters and a number of constitutively expressed and inducible genes. This work illustrates how TARGATT™ and Ansa Biotechnologies’ enzymatic DNA synthesis platform together enable rapid, efficient integration of large, complex genetic payloads — accelerating cell engineering timelines from months to weeks.
Methods
Cell culture and maintenance: The TARGATT™ hiPSC Master Cell Line (AST-9450), which contains a TARGATT™ landing pad at the H11 locus, was cultured according to the manufacturer’s instructions, passaging every 4-6 days.
Donor plasmid construction: The 60 kb construct (50 kb payload + 10 kb backbone) was synthesized by Ansa Biotechnologies using its enzymatic DNA synthesis platform. The sequence was divided into 80 segments of 600 bp – 700 bp that were synthesized as Ansamer™ contiguous oligonucleotides and amplified using PCR. The amplicons were cloned using E. coli and hierarchically assembled through two subsequent stages into the final full-length construct with the Ansa pStable-Ind-Chlor inducible-copy BAC backbone.
Transfection and integration: TARGATT™ hiPSC cells were transfected using protocols optimized for efficiency in iPSCs. Briefly, 1×10⁶ cells per well of a 6-well plate were co-transfected with 5.5 μg of Ansa’s 50 kb donor plasmid and 1.4 μg of TARGATT™ integrase plasmid. Cells were cultured for 48 hours post-transfection.
Selection and enrichment: Three days post-transfection, Zeocin selection (1.5 µg/mL) was initiated and maintained for one week. Surviving cells were expanded and analyzed for integration efficiency using flow cytometry.
Molecular and functional characterization: Genomic DNA was extracted and junction PCRs were performed using defined primers spanning the 5′ and 3′ integration sites. Surface expression of a constitutively expressed immune evasion protein was assessed by immunofluorescence flow cytometry. To evaluate inducible expression, cell pools were treated with doxycycline for 72 hours, stained with HaloTag TMR ligand to label the HaloTag, and fluorescence was measured using flow cytometry.
Results
To demonstrate targeted knock-in of a 50 kb construct into hiPSC cells, we designed a multi-functional payload containing four expression cassettes: cassette 1 constitutively expresses an immune evasion gene; cassette 2 constitutively expresses two immune evasion genes, Zeocin for selection, and the Tet-On® 3G transactivator; cassette 3 constitutively expresses a therapeutic payload and a miniGFP gene; and cassette 4 inducibly expresses a genome engineering gene and a HaloTag. Each expression cassette includes regulatory and co-expression elements such as Kozak sequences, WPREs, IRES, and polyA tails (Figure 1). The construct incorporates insulator elements between expression cassettes to prevent transcriptional interference.
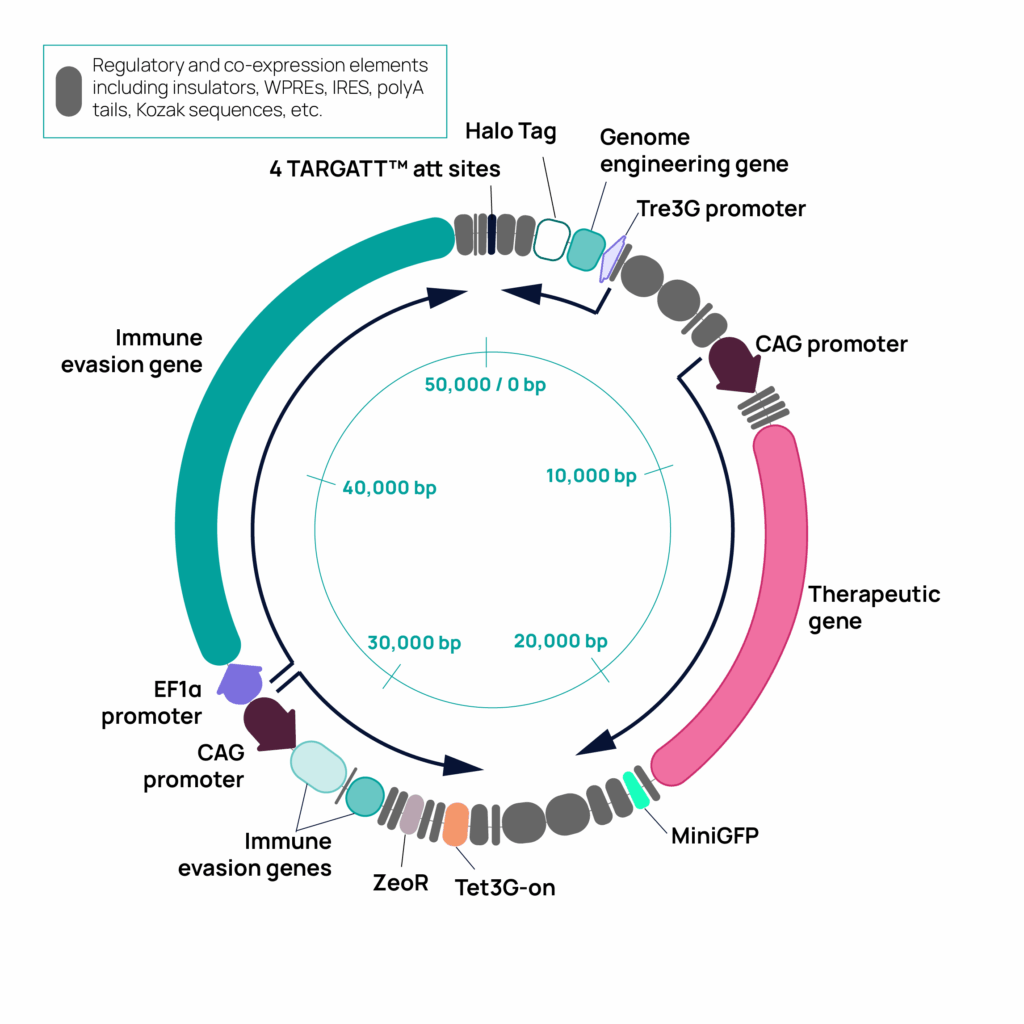
Figure 1. The 50 kb construct includes four expression cassettes with multiple functional elements. Expression cassettes are indicated by the curbed arrows, with key elements in color, as indicated, and elements to support robust gene and protein expression in gray.
Ansa Biotechnologies synthesized this 50 kb construct in less than one month using its enzymatic DNA synthesis platform. The construct was divided into 80 segments 600 bp – 700 bp long that were synthesized as individual contiguous Ansamer oligonucleotides up to 760 nt in length (including adaptors) that were cloned and then recombined into the full-length 50 kb insert through two subsequent rounds of assembly using a proprietary recombination method (Figure 2).
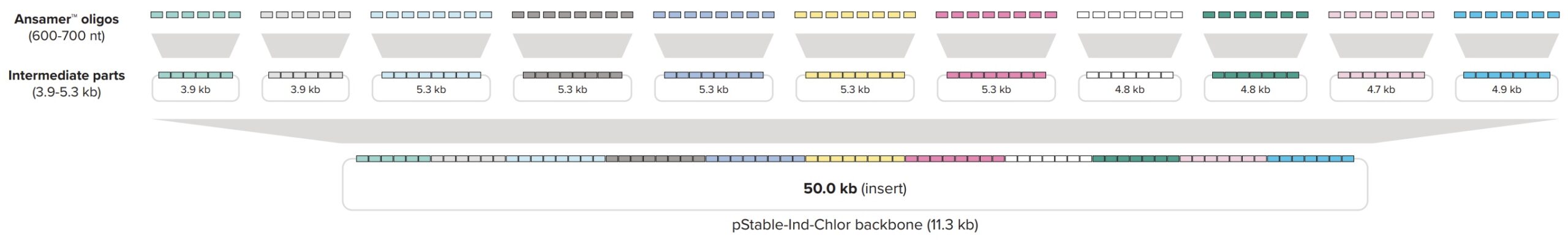
Figure 2. Hierarchical assembly strategy using 80 Ansamer oligonucleotides assembled through 11 subassemblies into the final construct.
The sequence-agnostic synthesis of long Ansamers enabled the construction of complex features within the construct, including repeats, hairpins, and regions of skewed GC content. The insert was cloned into Ansa’s preferred backbone for large constructs, pStable-Ind-Chlor, a single-copy vector that can be induced to mid-copy number using arabinose. This backbone facilitates stable propagation of large plasmids while enabling production of micrograms of DNA from a 2 mL liquid culture miniprep.
Following co-transfection of the donor construct with the TARGATT™ integrase plasmid, we observed ~88% integration efficiency after Zeocin selection, as determined by PCR of 34 clonal isolates. Junction PCR confirmed site-specific integration at the H11 locus, with both 5′ and 3′ junctions showing the expected band sizes of 1122 bp and 513 bp respectively (data not shown). Flow cytometry analysis showed robust expression of one of the immune evasion proteins, confirming functional expression of cassette 2 (Figure 3).
To validate inducible expression of cassette 4, we treated the donor-integrated cells with doxycycline for 72 hours and stained for HaloTag expression. Using flow cytometry, we observed that 23% of doxycycline-treated cells expressed the HaloTag versus negligible expression in untreated control cells (Figure 4).
Figure 4. Donor-integrated cells show inducible HaloTag expression. (A) Uninduced cells show negligible HaloTag expression. (B) After induction with doxycycline for 72 hrs, 23% of cells show HaloTag expression. The number of cells expressing the HaloTag increased over time (data not shown), consistent with robust inducibility.
Find out how your lab can leverage TARGATT™ technology with custom cell line knock-in services and iPSC, HEK293, and CHO kits.
Discussion
This work establishes that TARGATT™ technology can efficiently integrate constructs as large as 50 kb into human iPSCs, representing a 2.5-fold increase over previously characterized payloads⁴.
The rapid synthesis timeline achieved by Ansa’s enzymatic platform addresses a critical bottleneck in cell line engineering. Traditional approaches require 2+ months to build constructs of this size and often experience unpredictable delays, typically associated with regions of the sequence containing repeats or high GC content⁵. In fact, Applied Stem Cell estimates that using Ansa’s service to obtain even a 20 kb construct would save at least one month and 38% of the cost (time and materials) compared to building such a construct in-house. The company had not previously attempted to build 50 kb constructs in house because the associated costs and timelines were so unpredictable. By contrast, the one-month turnaround time of Ansa’s long DNA synthesis service dramatically accelerates iterative design cycles essential for optimizing complex genetic modifications.
The 50 kb payload capacity demonstrated here significantly expands the applications for site-specific integration. Screening libraries can now incorporate full-length proteins with native regulatory elements rather than truncated variants. For protein expression applications, entire multi-protein complexes with defined stoichiometry, such as full-length IgGs or heterodimeric cytokines, can easily be introduced in a single integration event.
Gene therapy applications benefit from the ability to create stable packaging lines expressing all required viral components. In cell therapy applications, sophisticated hypoimmunogenic designs can be easily implemented as well as complex CAR constructs that incorporate multiple signaling domains and logic gates. Disease modeling particularly benefits, as entire gene loci including introns and regulatory regions can be humanized to create more physiologically relevant models⁶.
Conclusion
The combination of Ansa’s rapid synthesis of ultra-long DNA constructs with ASC’s TARGATT™ large knock-in technology addresses two critical bottlenecks in advanced mammalian cell line engineering.
By demonstrating successful 50 kb integration with multi-functional genetic elements, this work expands the toolkit available for creating sophisticated cellular models and engineered cell lines. As genome engineering for screening, protein expression, gene therapy, cell therapy, and disease modeling grows increasingly complex, the ability to rapidly build and integrate large DNA constructs is essential for both research and therapeutic development.
The Tet-On® inducible expression system referenced herein is utilized by ASC strictly for internal research purposes. Such use is not intended for commercial distribution, sale, or transfer to third parties.
References
- Adli M. The CRISPR tool kit for genome editing and beyond. Nat Commun. 2018;9:1911.
- Duportet X, et al. A platform for rapid prototyping of synthetic gene networks in mammalian cells. Nucleic Acids Res. 2014;42(21):13440-13451.
- Fong J, et al. Transgene integration in mammalian cells: The tools, the challenges, and the future. Cell Systems. 2025; 101426.
- Chi X, et al. A system for site-specific integration of transgenes in mammalian cells. PLoS One. 2019;14(7):e0219842.
- Kosuri S, Church GM. Large-scale de novo DNA synthesis: technologies and applications. Nat Methods. 2014;11(5):499-507.
- Soldner F, Jaenisch R. iPSC disease modeling. Science. 2012;338(6111):1155-1156.
Ready to learn more?
Fill out the contact form below and a team member will be in touch within one business day.