Solving the DNA knock-in problem
Enabling technology for a new generation of cell therapy development
Cell therapy has emerged as a transformative therapeutic modality, capable of addressing diseases that are otherwise intractable with small molecules or biologics. Clinical success, particularly with CAR-T therapies in oncology, has validated the paradigm [1-3]. However, broader expansion into solid tumors, autoimmune disease, and regenerative medicine has been slower than anticipated. The central barrier is not simply immune rejection, but the complexity of engineering cells that can both evade immune detection and sustain therapeutic function over time [4, 5].
At the core of this challenge lies a fundamental technical bottleneck—the DNA knock-in problem.
Why engineering is a barrier to durable cell therapy
To achieve robust engraftment, allogeneic cell therapies must evade multiple arms of the immune system simultaneously. This requires coordinated genetic modification—disrupting antigen presentation pathways (e.g., B2M and CIITA knock-out), suppressing innate immune responses (e.g., CD47 overexpression), and mitigating NK cell activation through expression of non-classical HLA molecules. These pathways are interdependent; incomplete engineering results in rapid immune-mediated rejection [4, 6].
At the same time, many of the most promising applications of cell therapy—particularly those addressing loss of cellular function—require the precise addition of therapeutic payloads. These may include full-length genes to restore function, synthetic receptors such as CARs, or multi-gene constructs designed to enhance persistence and activity. In other words, developers must not only knock genes out, but also reliably knock genes in, often at significant size and complexity.
Knock-ins are now the bottleneck
This dual requirement—multiplex knock-out for immune evasion and large, precise knock-in for therapeutic function—creates a compounding engineering challenge. While knock-out technologies have advanced rapidly, as can be seen in the rapid evolution of CRISPR systems, knock-in remains a major bottleneck. Conventional approaches such as viral vectors or transposon systems introduce payloads in a semi-random manner, leading to variable expression, potential silencing, and safety concerns related to insertional mutagenesis [7, 8]. Just as importantly, these approaches are poorly suited for large or multi-gene constructs and lack the precision required for next-generation therapies.
As a result, developers face a paradox—the biology of durable cell therapy demands increasingly sophisticated genetic programs, but the tools available to install those programs had been limiting.
This challenge is particularly evident in emerging allogeneic platforms such as induced pluripotent stem cells (iPSCs). These systems offer a scalable, universal starting material that can be engineered once and deployed across multiple indications. However, realizing this vision depends on the ability to introduce complex genetic payloads in a precise, reproducible, and stable manner—without compromising cell fitness or safety.
This is where site-specific technologies that can handle large DNA fragments become essential.
A shift toward site-specific, large-fragment knock-in
The TARGATT™ platform addresses the knock-in problem by enabling site-specific insertion of DNA payloadsup to 50 kb at a proven safe harbor site (Figure 1).
By avoiding random integration and inserting into a well-characterized safe-harbor site, the TARGATT™ platform enables consistent and predictable gene expression—critical for both safety and efficacy. Its large cargo capacity allows developers to introduce multi-gene immune evasion cassettes, therapeutic transgenes, and regulatory elements in a single edit, simplifying what is otherwise a fragmented engineering process.
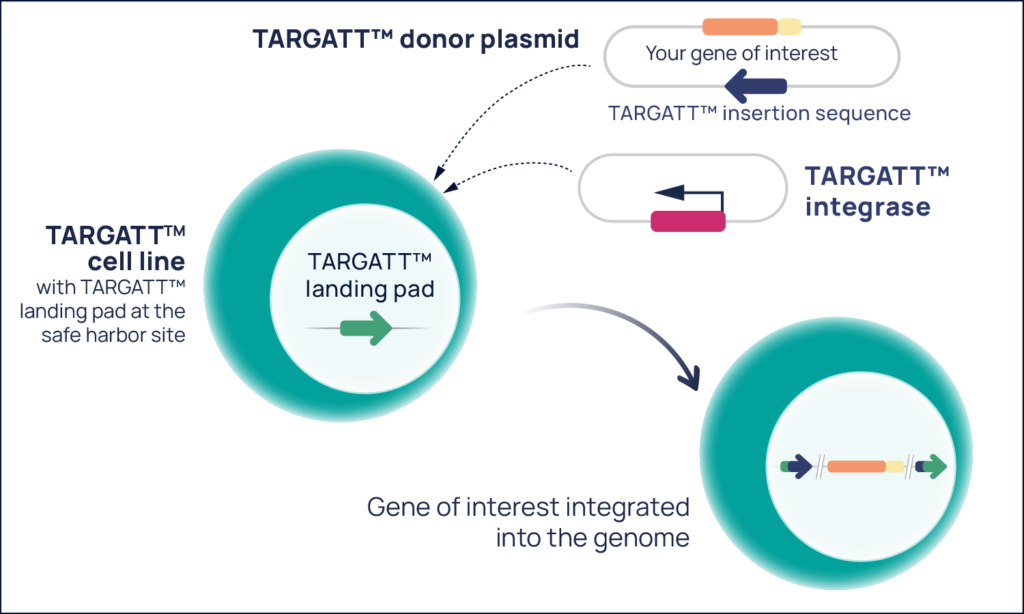
Figure 1. The TARGATT™ platform consists of a donor plasmid with your gene of interest, a plasmid that expresses the TARGATT™ integrase, and a cell line that contains the TARGATT™ landing pad at the proven H11 safe harbor site—learn more.
Importantly, TARGATT™ technology complements existing genome editing strategies. Knock-out edits can eliminate immunogenic pathways while TARGATT™ installs the complex genetic programs required for immune evasion and therapeutic functions. Together, these approaches enable a new level of control over cell design.
Solving the DNA knock-in problem
As the field advances toward durable, off-the-shelf therapies, the demands on genetic engineering will only increase. Success will depend not just on removing problematic genes, but on the ability to add sophisticated functionality in a controlled and scalable way.
In this context, the knock-in problem is no longer secondary—it is a defining constraint on the future of cell therapy. Technologies that solve this problem, like the TARGATT™ platform, are more than incremental improvements—they are foundational enablers of a new generation of cell-based medicines.
Find out how your lab can solve the DNA knock-in problem with TARGATT™ technology.
References
- Park JH, Rivière I, Gonen M, et al. Long-Term Follow-up of CD19 CAR Therapy in Acute Lymphoblastic Leukemia. N Engl J Med. 2018;378(5):449-459. doi:10.1056/NEJMoa1709919
- June CH, Sadelain M. Chimeric Antigen Receptor Therapy. N Engl J Med. 2018;379(1):64-73. doi:10.1056/NEJMra1706169
- Maude SL, Laetsch TW, Buechner J, et al. Tisagenlecleucel in Children and Young Adults with B-Cell Lymphoblastic Leukemia. N Engl J Med. 2018;378(5):439-448. doi:10.1056/NEJMoa1709866
- Deuse T, Schrepfer S. Progress and challenges in developing allogeneic cell therapies. Cell Stem Cell. 2025;32(4):513-528. doi:10.1016/j.stem.2025.03.004
- Shapiro AMJ, Lakey JRT, Ryan EA, et al. Islet Transplantation in Seven Patients with Type 1 Diabetes Mellitus Using a Glucocorticoid-Free Immunosuppressive Regimen. N Engl J Med. 2000;343(4):230-238. doi:10.1056/NEJM200007273430401
- Otsuka R, Wada H, Murata T, Seino K ichiro. Immune reaction and regulation in transplantation based on pluripotent stem cell technology. Inflamm Regen. 2020;40(1):12. doi:10.1186/s41232-020-00125-8
- Hacein-Bey-Abina S, Von Kalle C, Schmidt M, et al. LMO2-Associated Clonal T Cell Proliferation in Two Patients after Gene Therapy for SCID-X1. Science. 2003;302(5644):415-419. doi:10.1126/science.1088547
- Hacein-Bey-Abina S, Kalle C von, Schmidt M, et al. A Serious Adverse Event after Successful Gene Therapy for X-Linked Severe Combined Immunodeficiency. N Engl J Med. 2003;348(3):255-256. doi:10.1056/NEJM200301163480314